Services
For more information on the Systems Biology module or to request services, please contact Anthony St. Leger at anthony.stleger@pitt.edu.
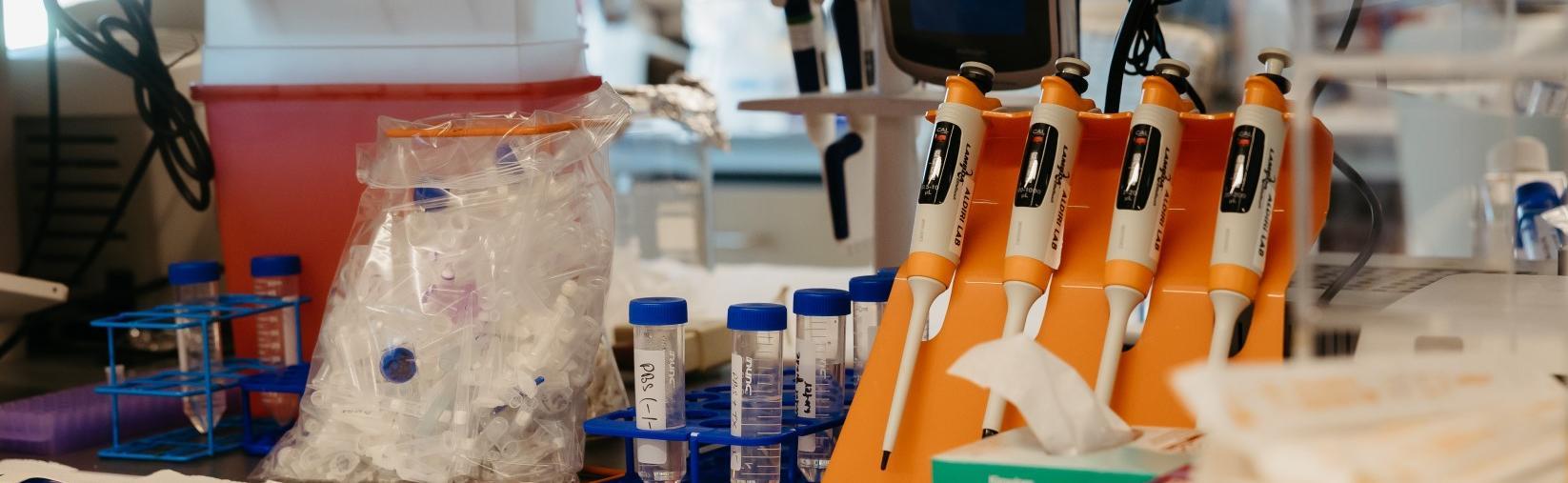
The Systems Biology Core at the University of Pittsburgh Department of Ophthalmology is a cutting-edge facility focused on flow cytometry and fluorescence-activated cell sorting (FACS)-based cell analysis and isolation. We also help support single-cell analysis of bulk and sorted populations of cells from various species that include mouse, zebrafish, nonhuman primate, human, and others.
Analysis includes descriptions of the cell (size, granularity, protein expression), isolating cells to express proteins or create clones, and sorting cells into various populations of interest. Similarly, we can qualitatively measure fluorescent protein expression for analysis and/or cell isolation. The equipment in the System Biology module can conduct analysis on both a single cell and population cell basis.
Backed by daily quality control and expert staff support, the core is a vital resource for advancing ophthalmology research at the University of Pittsburgh.
Equipment
Our hallmark cell analyzers are the Beckman Coulter Cytoflex LX and Sony ID7000. Both pieces of equipment are powerful tools for phenotypic characterization and can analyze up to 14 and 55 parameters, respectively. The Sony MA900 and MACS Tyto provide complementary cell sorting capabilities, with the former sorting up to four populations at low pressure, and the latter using a gravity-based approach for low-pressure sorts and maximum cell viability. These instruments support a range of applications, from detailed cell profiling to the isolation of cells for advanced molecular analyses like RNAseq and chromatin studies.
Downstream of cell sorting, the core is equipped with instruments that can assist users in the preparation of single-cell RNAseq/ATACseq libraries and multiome library prep. Those instruments include a cell counter, tapestation, and 10X Chromium machine.
Project Details
For each project, Dr. St Leger will consult with the researcher to define their end goal and design an experimental plan to accomplish that goal. All researchers can download analytical software from each machine and install it on their PC. Licenses are also available to borrow for more sophisticated software that can compress multiple parameters into a single figure.
An example of a scatter plot produced by the module’s equipment is below. The x-axis is FSC (forward scatter), showing how big a cell is, while the y-axis is SSC (side scatter), showing granularity.
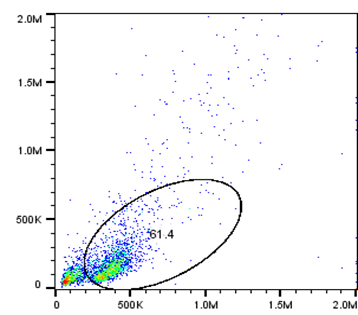
Additionally, an example of cell sorting output is shown below:
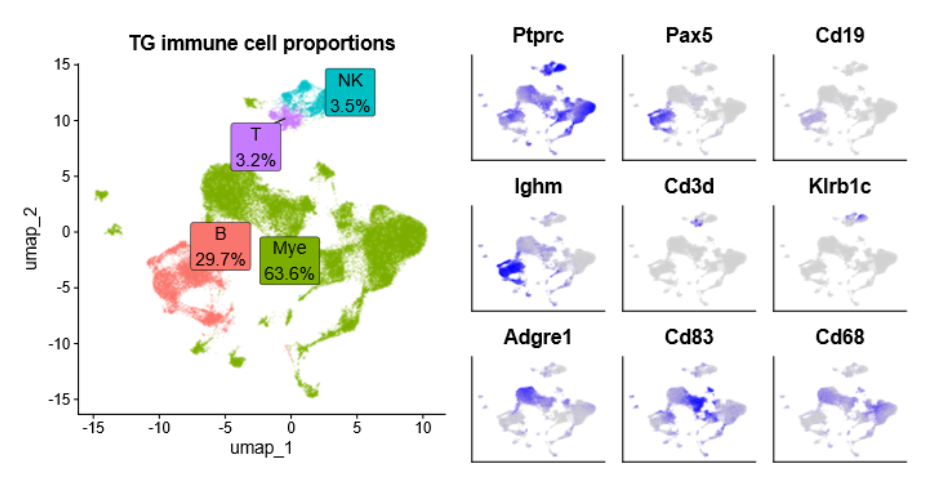
In this example, immune cells are sorted in preparation for single-cell RNAseq analysis. Cell sorting can also be used to create profiles of any cells and tissues of interest, including immune receptors and autoimmune cells. The cell sorter is located inside a biosafety cabinet, so infected tissues and viruses are contained.
Other examples of projects undertaken by the System Biology module include comparing cells to one another with relative analysis, describing population shifts, and isolating cells, bacteria, or fungi to create clones.